Lignin depolymerization by fungal secretomes and a microbial sink
Abstract
Fungal ligninolytic enzymes are able to depolymerize solid lignin and the presence of an aromatic catabolic bacterium enhances this effect.
- Authors:
-
- National Renewable Energy Lab. (NREL), Golden, CO (United States)
- Pacific Northwest National Lab. (PNNL), Richland, WA (United States)
- Consejo Superior de Investigaciones CientÃficas (CSIC), Madrid (Spain)
- Joint BioEnergy Institute (JBEI), Emeryville, CA (United States); Lawrence Berkeley National Lab. (LBNL), Berkeley, CA (United States)
- Joint BioEnergy Institute (JBEI), Emeryville, CA (United States); Sandia National Lab. (SNL-CA), Livermore, CA (United States)
- Publication Date:
- Research Org.:
- National Renewable Energy Laboratory (NREL), Golden, CO (United States); Lawrence Berkeley National Laboratory (LBNL), Berkeley, CA (United States)
- Sponsoring Org.:
- USDOE Office of Energy Efficiency and Renewable Energy (EERE), Sustainable Transportation Office. Bioenergy Technologies Office (BETO); USDOE Office of Science (SC), Biological and Environmental Research (BER)
- OSTI Identifier:
- 1331971
- Alternate Identifier(s):
- OSTI ID: 1440916
- Report Number(s):
- NREL/JA-5100-67108
Journal ID: ISSN 1463-9262
- Grant/Contract Number:
- AC36-08GO28308; AC02-05CH11231
- Resource Type:
- Accepted Manuscript
- Journal Name:
- Green Chemistry
- Additional Journal Information:
- Journal Volume: 18; Journal Issue: 22; Journal ID: ISSN 1463-9262
- Publisher:
- Royal Society of Chemistry
- Country of Publication:
- United States
- Language:
- English
- Subject:
- 09 BIOMASS FUELS; 37 INORGANIC, ORGANIC, PHYSICAL, AND ANALYTICAL CHEMISTRY; lignin depolymerization; fungal secretomes; biorefinery
Citation Formats
Salvachua, Davinia, Katahira, Rui, Cleveland, Nicholas S., Khanna, Payal, Resch, Michael G., Black, Brenna A., Purvine, Samuel O., Zink, Erika M., Prieto, Alicia, Martinez, Maria J., Martinez, Angel T., Simmons, Blake A., Gladden, John M., and Beckham, Gregg T. Lignin depolymerization by fungal secretomes and a microbial sink. United States: N. p., 2016.
Web. doi:10.1039/C6GC01531J.
Salvachua, Davinia, Katahira, Rui, Cleveland, Nicholas S., Khanna, Payal, Resch, Michael G., Black, Brenna A., Purvine, Samuel O., Zink, Erika M., Prieto, Alicia, Martinez, Maria J., Martinez, Angel T., Simmons, Blake A., Gladden, John M., & Beckham, Gregg T. Lignin depolymerization by fungal secretomes and a microbial sink. United States. https://doi.org/10.1039/C6GC01531J
Salvachua, Davinia, Katahira, Rui, Cleveland, Nicholas S., Khanna, Payal, Resch, Michael G., Black, Brenna A., Purvine, Samuel O., Zink, Erika M., Prieto, Alicia, Martinez, Maria J., Martinez, Angel T., Simmons, Blake A., Gladden, John M., and Beckham, Gregg T. Thu .
"Lignin depolymerization by fungal secretomes and a microbial sink". United States. https://doi.org/10.1039/C6GC01531J. https://www.osti.gov/servlets/purl/1331971.
@article{osti_1331971,
title = {Lignin depolymerization by fungal secretomes and a microbial sink},
author = {Salvachua, Davinia and Katahira, Rui and Cleveland, Nicholas S. and Khanna, Payal and Resch, Michael G. and Black, Brenna A. and Purvine, Samuel O. and Zink, Erika M. and Prieto, Alicia and Martinez, Maria J. and Martinez, Angel T. and Simmons, Blake A. and Gladden, John M. and Beckham, Gregg T.},
abstractNote = {Fungal ligninolytic enzymes are able to depolymerize solid lignin and the presence of an aromatic catabolic bacterium enhances this effect.},
doi = {10.1039/C6GC01531J},
journal = {Green Chemistry},
number = 22,
volume = 18,
place = {United States},
year = {Thu Aug 25 00:00:00 EDT 2016},
month = {Thu Aug 25 00:00:00 EDT 2016}
}
Free Publicly Available Full Text
Publisher's Version of Record
Other availability
Cited by: 69 works
Citation information provided by
Web of Science
Web of Science
Figures / Tables:
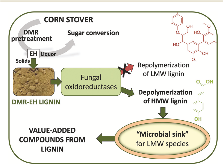 Fig. 1: Microbial sink concept. The presence of bacteria with ligninolytic enzymes, which are able to depolymerize high molecular weight (HMW) lignin, may act as a sink to avoid repolymerization of low molecular weight (LMW) aromatic compounds and enhance biological lignin depolymerization.
Fig. 1: Microbial sink concept. The presence of bacteria with ligninolytic enzymes, which are able to depolymerize high molecular weight (HMW) lignin, may act as a sink to avoid repolymerization of low molecular weight (LMW) aromatic compounds and enhance biological lignin depolymerization.
All figures and tables
(10 total)
Save to My Library
You must Sign In or Create an Account in order to save documents to your library.
Works referenced in this record:
Cell wall modifications triggered by the down-regulation of Coumarate 3-hydroxylase-1 in maize
journal, July 2015
- Fornalé, Silvia; Rencoret, Jorge; Garcia-Calvo, Laura
- Plant Science, Vol. 236
Laccase detoxification of steam-exploded wheat straw for second generation bioethanol
journal, December 2009
- Jurado, Miguel; Prieto, Alicia; Martínez-Alcalá, Ángeles
- Bioresource Technology, Vol. 100, Issue 24
Fungal Laccases and Their Applications in Bioremediation
journal, January 2014
- Viswanath, Buddolla; Rajesh, Bandi; Janardhan, Avilala
- Enzyme Research, Vol. 2014
Fungal Cellulases
journal, January 2015
- Payne, Christina M.; Knott, Brandon C.; Mayes, Heather B.
- Chemical Reviews, Vol. 115, Issue 3
Lignin Changes after Steam Explosion and Laccase-Mediator Treatment of Eucalyptus Wood Chips
journal, August 2011
- Martin-Sampedro, Raquel; Capanema, Ewellyn A.; Hoeger, Ingrid
- Journal of Agricultural and Food Chemistry, Vol. 59, Issue 16
Towards lignin consolidated bioprocessing: simultaneous lignin depolymerization and product generation by bacteria
journal, January 2015
- Salvachúa, Davinia; Karp, Eric M.; Nimlos, Claire T.
- Green Chemistry, Vol. 17, Issue 11
Sugar recoveries from wheat straw following treatments with the fungus Irpex lacteus
journal, March 2013
- Salvachúa, Davinia; Prieto, Alicia; Vaquero, María Eugenia
- Bioresource Technology, Vol. 131
Identification of DypB from Rhodococcus jostii RHA1 as a Lignin Peroxidase
journal, June 2011
- Ahmad, Mark; Roberts, Joseph N.; Hardiman, Elizabeth M.
- Biochemistry, Vol. 50, Issue 23
Upgrading of an industrial lignin by using laccase produced by Fusarium proliferatum and different laccase-mediator systems
journal, January 2006
- Hernández Fernaud, J. R.; Carnicero, A.; Perestelo, F.
- Enzyme and Microbial Technology, Vol. 38, Issue 1-2
Oxidative upgrade of lignin – Recent routes reviewed
journal, June 2013
- Lange, Heiko; Decina, Silvia; Crestini, Claudia
- European Polymer Journal, Vol. 49, Issue 6
Specific degradation ofβ-aryl ether linkage in synthetic lignin (dehydrogenative polymerizate) by bacterial enzymes ofSphingomonas paucimobilis SYK-6 produced in recombinantEscherichia coli
journal, October 2002
- Sonoki, Tomonori; Iimura, Yosuke; Masai, Eiji
- Journal of Wood Science, Vol. 48, Issue 5
Efficient bleaching of non-wood high-quality paper pulp using laccase-mediator system
journal, August 2004
- Camarero, Susana; Garcı́a, Olga; Vidal, Teresa
- Enzyme and Microbial Technology, Vol. 35, Issue 2-3
Purification and characterization of peroxidases from the dye-decolorizing fungus Bjerkandera adusta
journal, August 1998
- Heinfling, Annette; MartıÌnez, MarıÌa Jesús; MartıÌnez, Angel T.
- FEMS Microbiology Letters, Vol. 165, Issue 1
Ligninolytic peroxidase genes in the oyster mushroom genome: heterologous expression, molecular structure, catalytic and stability properties, and lignin-degrading ability
journal, January 2014
- Fernández-Fueyo, Elena; Ruiz-Dueñas, Francisco J.; Martínez, María
- Biotechnology for Biofuels, Vol. 7, Issue 1
Helically agitated mixing in dry dilute acid pretreatment enhances the bioconversion of corn stover into ethanol
journal, January 2014
- He, Yanqing; Zhang, Longping; Zhang, Jian
- Biotechnology for Biofuels, Vol. 7, Issue 1
DeconMSn: a software tool for accurate parent ion monoisotopic mass determination for tandem mass spectra
journal, February 2008
- Mayampurath, A. M.; Jaitly, N.; Purvine, S. O.
- Bioinformatics, Vol. 24, Issue 7
Pretreatment with laccase and a phenolic mediator degrades lignin and enhances saccharification of Eucalyptus feedstock
journal, January 2014
- Rico, Alejandro; Rencoret, Jorge; del Río, José C.
- Biotechnology for Biofuels, Vol. 7, Issue 1
A method for the quantitative recovery of protein in dilute solution in the presence of detergents and lipids
journal, April 1984
- Wessel, D.; Flügge, U. I.
- Analytical Biochemistry, Vol. 138, Issue 1
Application of the Accurate Mass and Time Tag Approach to the Proteome Analysis of Sub-cellular Fractions Obtained from Rhodobacter s phaeroides 2.4.1. Aerobic and Photosynthetic Cell Cultures
journal, August 2006
- Callister, Stephen J.; Dominguez, Miguel A.; Nicora, Carrie D.
- Journal of Proteome Research, Vol. 5, Issue 8
SignalP 4.0: discriminating signal peptides from transmembrane regions
journal, September 2011
- Petersen, Thomas Nordahl; Brunak, Søren; von Heijne, Gunnar
- Nature Methods, Vol. 8, Issue 10
Bacterial versus fungal laccase: potential for micropollutant degradation
journal, January 2013
- Margot, Jonas; Bennati-Granier, Chloé; Maillard, Julien
- AMB Express, Vol. 3, Issue 1
Chemically Etched Open Tubular and Monolithic Emitters for Nanoelectrospray Ionization Mass Spectrometry
journal, November 2006
- Kelly, Ryan T.; Page, Jason S.; Luo, Quanzhou
- Analytical Chemistry, Vol. 78, Issue 22, p. 7796-7801
Differential proteomic analysis of the secretome of Irpex lacteus and other white-rot fungi during wheat straw pretreatment
journal, January 2013
- Salvachúa, Davinia; Martínez, Angel T.; Tien, Ming
- Biotechnology for Biofuels, Vol. 6, Issue 1
Paper pulp delignification using laccase and natural mediators
journal, April 2007
- Camarero, Susana; Ibarra, David; Martínez, Ángel T.
- Enzyme and Microbial Technology, Vol. 40, Issue 5
Solution-state 2D NMR of Ball-milled Plant Cell Wall Gels in DMSO-d 6
journal, March 2008
- Kim, Hoon; Ralph, John; Akiyama, Takuya
- BioEnergy Research, Vol. 1, Issue 1, p. 56-66
Characterization of a Novel Dye-Decolorizing Peroxidase (DyP)-Type Enzyme from Irpex lacteus and Its Application in Enzymatic Hydrolysis of Wheat Straw
journal, May 2013
- Salvachua, D.; Prieto, A.; Martinez, A. T.
- Applied and Environmental Microbiology, Vol. 79, Issue 14
Laccases and their natural mediators: Biotechnological tools for sustainable eco-friendly processes
journal, November 2010
- Cañas, Ana I.; Camarero, Susana
- Biotechnology Advances, Vol. 28, Issue 6
Lignin valorization through integrated biological funneling and chemical catalysis
journal, August 2014
- Linger, J. G.; Vardon, D. R.; Guarnieri, M. T.
- Proceedings of the National Academy of Sciences, Vol. 111, Issue 33, p. 12013-12018
Enzymatic and Fungal Treatments on Sugarcane Bagasse for the Production of Mechanical Pulps
journal, July 2004
- Ramos, Juan; Rojas, Teresa; Navarro, Fernando
- Journal of Agricultural and Food Chemistry, Vol. 52, Issue 16
Basidiomycete DyPs: Genomic diversity, structural–functional aspects, reaction mechanism and environmental significance
journal, May 2015
- Linde, Dolores; Ruiz-Dueñas, Francisco J.; Fernández-Fueyo, Elena
- Archives of Biochemistry and Biophysics, Vol. 574
Fungal aryl-alcohol oxidase: a peroxide-producing flavoenzyme involved in lignin degradation
journal, January 2012
- Hernández-Ortega, Aitor; Ferreira, Patricia; Martínez, Angel T.
- Applied Microbiology and Biotechnology, Vol. 93, Issue 4
The emerging role for bacteria in lignin degradation and bio-product formation
journal, June 2011
- Bugg, Timothy DH; Ahmad, Mark; Hardiman, Elizabeth M.
- Current Opinion in Biotechnology, Vol. 22, Issue 3
A secretomic view of woody and nonwoody lignocellulose degradation by Pleurotus ostreatus
journal, February 2016
- Fernández-Fueyo, Elena; Ruiz-Dueñas, Francisco J.; López-Lucendo, María F.
- Biotechnology for Biofuels, Vol. 9, Issue 1
Lignin Valorization: Improving Lignin Processing in the Biorefinery
journal, May 2014
- Ragauskas, A. J.; Beckham, G. T.; Biddy, M. J.
- Science, Vol. 344, Issue 6185, p. 1246843-1246843
A Mechanistic Investigation of Acid-Catalyzed Cleavage of Aryl-Ether Linkages: Implications for Lignin Depolymerization in Acidic Environments
journal, November 2013
- Sturgeon, Matthew R.; Kim, Seonah; Lawrence, Kelsey
- ACS Sustainable Chemistry & Engineering, Vol. 2, Issue 3, p. 472-485
Demonstration of laccase-based removal of lignin from wood and non-wood plant feedstocks
journal, September 2012
- Gutiérrez, Ana; Rencoret, Jorge; Cadena, Edith M.
- Bioresource Technology, Vol. 119
Improved ethanol yield and reduced minimum ethanol selling price (MESP) by modifying low severity dilute acid pretreatment with deacetylation and mechanical refining: 2) Techno-economic analysis
journal, January 2012
- Tao, Ling; Chen, Xiaowen; Aden, Andy
- Biotechnology for Biofuels, Vol. 5, Issue 1
Peroxidases depolymerize lignin in organic media but not in water
journal, September 1986
- Dordick, J. S.; Marletta, M. A.; Klibanov, A. M.
- Proceedings of the National Academy of Sciences, Vol. 83, Issue 17
Pathways for degradation of lignin in bacteria and fungi
journal, January 2011
- Bugg, Timothy D. H.; Ahmad, Mark; Hardiman, Elizabeth M.
- Natural Product Reports, Vol. 28, Issue 12, p. 1883-1896
New insights into the structure and composition of technical lignins: a comparative characterisation study
journal, January 2016
- Constant, Sandra; Wienk, Hans L. J.; Frissen, Augustinus E.
- Green Chemistry, Vol. 18, Issue 9
Demonstration of Lignin-to-Peroxidase Direct Electron Transfer: A TRANSIENT-STATE KINETICS, DIRECTED MUTAGENESIS, EPR, AND NMR STUDY
journal, August 2015
- Sáez-Jiménez, Verónica; Baratto, Maria Camilla; Pogni, Rebecca
- Journal of Biological Chemistry, Vol. 290, Issue 38
Structural modification of eucalypt pulp lignin in a totally chlorine-free bleaching sequence including a laccase-mediator stage
journal, November 2007
- Ibarra, David; Chávez, María Isabel; Rencoret, Jorge
- Holzforschung, Vol. 61, Issue 6
Veratryl Alcohol Oxidase from Pleurotus ostreatus Participates in Lignin Biodegradation and Prevents Polymerization of Laccase-oxidized Substrates
journal, February 1995
- Marzullo, Liberato; Cannio, Raffaele; Giardina, Paola
- Journal of Biological Chemistry, Vol. 270, Issue 8
Whole plant cell wall characterization using solution-state 2D NMR
journal, August 2012
- Mansfield, Shawn D.; Kim, Hoon; Lu, Fachuang
- Nature Protocols, Vol. 7, Issue 9
Versatile peroxidase as a valuable tool for generating new biomolecules by homogeneous and heterogeneous cross-linking
journal, May 2013
- Salvachúa, Davinia; Prieto, Alicia; Mattinen, Maija-Liisa
- Enzyme and Microbial Technology, Vol. 52, Issue 6-7
Kinetic analysis of regional agro-industrial waste combustion
journal, July 2016
- Fernandez, Anabel; Saffe, Alejandra; Mazza, Germán
- Biofuels, Vol. 8, Issue 1
Opportunities and challenges in biological lignin valorization
journal, December 2016
- Beckham, Gregg T.; Johnson, Christopher W.; Karp, Eric M.
- Current Opinion in Biotechnology, Vol. 42
Adipic acid production from lignin
journal, January 2015
- Vardon, Derek R.; Franden, Mary Ann; Johnson, Christopher W.
- Energy & Environmental Science, Vol. 8, Issue 2
Target-Decoy Search Strategy for Mass Spectrometry-Based Proteomics
book, December 2009
- Elias, Joshua E.; Gygi, Steven P.
- Methods in Molecular Biology
Fungal pretreatment: An alternative in second-generation ethanol from wheat straw
journal, August 2011
- Salvachúa, Davinia; Prieto, Alicia; López-Abelairas, María
- Bioresource Technology, Vol. 102, Issue 16
Basic local alignment search tool
journal, October 1990
- Altschul, Stephen F.; Gish, Warren; Miller, Webb
- Journal of Molecular Biology, Vol. 215, Issue 3, p. 403-410
DMR (deacetylation and mechanical refining) processing of corn stover achieves high monomeric sugar concentrations (230 g L −1 ) during enzymatic hydrolysis and high ethanol concentrations (>10% v/v) during fermentation without hydrolysate purification or concentration
journal, January 2016
- Chen, Xiaowen; Kuhn, Erik; Jennings, Edward W.
- Energy & Environmental Science, Vol. 9, Issue 4
Molecular biology and structure-function of lignin-degrading heme peroxidases
journal, April 2002
- Martı́nez, Angel T.
- Enzyme and Microbial Technology, Vol. 30, Issue 4
Synergistic enzymatic and microbial lignin conversion
journal, January 2016
- Zhao, Cheng; Xie, Shangxian; Pu, Yunqiao
- Green Chemistry, Vol. 18, Issue 5
From gene to biorefinery: microbial β-etherases as promising biocatalysts for lignin valorization
journal, September 2015
- Picart, Pere; de María, Pablo Domínguez; Schallmey, Anett
- Frontiers in Microbiology, Vol. 6
Effects of laccase on lignin depolymerization and enzymatic hydrolysis of ensiled corn stover
journal, August 2012
- Chen, Qin; Marshall, Megan N.; Geib, Scott M.
- Bioresource Technology, Vol. 117
Conversion of Milled Pine Wood by Manganese Peroxidase from Phlebia radiata
journal, October 2001
- Hofrichter, M.; Lundell, T.; Hatakka, A.
- Applied and Environmental Microbiology, Vol. 67, Issue 10
Biomass Fractionation for the Biorefinery: Heteronuclear Multiple Quantum Coherence–Nuclear Magnetic Resonance Investigation of Lignin Isolated from Solvent Fractionation of Switchgrass
journal, September 2011
- Bozell, Joseph J.; O'Lenick, C. J.; Warwick, Stacy
- Journal of Agricultural and Food Chemistry, Vol. 59, Issue 17
In vitro degradation of natural insoluble lignin in aqueous media by the extracellular peroxidases of Phanerochaete chrysosporium
journal, March 1998
- Thompson, David N.; Hames, Bonnie R.; Reddy, C. Adinarayana
- Biotechnology and Bioengineering, Vol. 57, Issue 6, p. 704-717
Novel multienzyme oxidative biocatalyst for lignin bioprocessing
journal, August 2011
- Crestini, Claudia; Melone, Federica; Saladino, Raffaele
- Bioorganic & Medicinal Chemistry, Vol. 19, Issue 16
p -Hydroxycinnamic Acids as Natural Mediators for Laccase Oxidation of Recalcitrant Compounds
journal, September 2008
- Camarero, Susana; CaÑas, Ana I.; Nousiainen, Paula
- Environmental Science & Technology, Vol. 42, Issue 17
Oxidation of milled wood lignin with laccase, tyrosinase and horseradish peroxidase
journal, December 2004
- Grönqvist, S.; Viikari, L.; Niku-Paavola, M. -L.
- Applied Microbiology and Biotechnology, Vol. 67, Issue 4
Improved Xylan Hydrolysis of Corn Stover by Deacetylation with High Solids Dilute Acid Pretreatment
journal, December 2011
- Chen, Xiaowen; Shekiro, Joseph; Elander, Rick
- Industrial & Engineering Chemistry Research, Vol. 51, Issue 1
Can laccases catalyze bond cleavage in lignin?
journal, January 2015
- Munk, Line; Sitarz, Anna K.; Kalyani, Dayanand C.
- Biotechnology Advances, Vol. 33, Issue 1
Towards industrially-feasible delignification and pitch removal by treating paper pulp with Myceliophthora thermophila laccase and a phenolic mediator
journal, June 2011
- Babot, Esteban D.; Rico, Alejandro; Rencoret, Jorge
- Bioresource Technology, Vol. 102, Issue 12
Self-made frits for nanoscale columns in proteomics
journal, October 2005
- Maiolica, Alessio; Borsotti, Dario; Rappsilber, Juri
- PROTEOMICS, Vol. 5, Issue 15
Fungal cellulases
journal, February 1992
- Wood, Thomas M.
- Biochemical Society Transactions, Vol. 20, Issue 1
Demonstration of lignin-to-peroxidase direct electron transfer. A TRANSIENT-STATE KINETICS, DIRECTED MUTAGENESIS, EPR AND NMR STUDY.
journal, December 2015
- Sáez-Jiménez, Verónica; Baratto, Maria Camilla; Pogni, Rebecca
- Journal of Biological Chemistry, Vol. 290, Issue 51
Works referencing / citing this record:
Biotransformation of lignin: Mechanisms, applications and future work
journal, October 2019
- Li, Xiang; Zheng, Yi
- Biotechnology Progress, Vol. 36, Issue 1
Bioethanol Mill Wastewater Purification by Combination of Coagulation-Flocculation and Microbial Treatment of Trametes versicolor INACC F200
journal, August 2019
- Sari, Ajeng Arum; Hadibarata, Tony; Hanifah, Ummu
- Water, Air, & Soil Pollution, Vol. 230, Issue 9
Lignin Depolymerization to BTXs
journal, September 2019
- Serrano, Luis; Cecilia, Juan Antonio; García-Sancho, Cristina
- Topics in Current Chemistry, Vol. 377, Issue 5
Enabling microbial syringol conversion through structure-guided protein engineering
journal, June 2019
- Machovina, Melodie M.; Mallinson, Sam J. B.; Knott, Brandon C.
- Proceedings of the National Academy of Sciences, Vol. 116, Issue 28
Metabolic engineering of Pseudomonas putida for increased polyhydroxyalkanoate production from lignin
journal, January 2020
- Salvachúa, Davinia; Rydzak, Thomas; Auwae, Raquel
- Microbial Biotechnology, Vol. 13, Issue 1
Enzyme biotechnology in degradation and modification of plant cell wall polymers
journal, August 2018
- Marjamaa, Kaisa; Kruus, Kristiina
- Physiologia Plantarum, Vol. 164, Issue 1
Development of a genetically programed vanillin-sensing bacterium for high-throughput screening of lignin-degrading enzyme libraries
journal, February 2017
- Sana, Barindra; Chia, Kuan Hui Burton; Raghavan, Sarada S.
- Biotechnology for Biofuels, Vol. 10, Issue 1
Differential β-glucosidase expression as a function of carbon source availability in Talaromyces amestolkiae: a genomic and proteomic approach
journal, June 2017
- de Eugenio, Laura I.; Méndez-Líter, Juan A.; Nieto-Domínguez, Manuel
- Biotechnology for Biofuels, Vol. 10, Issue 1
A bionic system with Fenton reaction and bacteria as a model for bioprocessing lignocellulosic biomass
journal, February 2018
- Zhang, Kejing; Si, Mengying; Liu, Dan
- Biotechnology for Biofuels, Vol. 11, Issue 1
Lignin Depolymerization to BTXs
book, September 2019
- Serrano, Luis; Cecilia, Juan Antonio; García‑Sancho, Cristina
- Topics in Current Chemistry Collections
Figures / Tables found in this record:
Figures/Tables have been extracted from DOE-funded journal article accepted manuscripts.

 Search WorldCat to find libraries that may hold this journal
Search WorldCat to find libraries that may hold this journal