Tracking microbial colonization in fecal microbiota transplantation experiments via genome-resolved metagenomics
Abstract
Fecal microbiota transplantation (FMT) is an effective treatment for recurrent Clostridium difficile infection and shows promise for treating other medical conditions associated with intestinal dysbioses. However, we lack a sufficient understanding of which microbial populations successfully colonize the recipient gut, and the widely used approaches to study the microbial ecology of FMT experiments fail to provide enough resolution to identify populations that are likely responsible for FMT-derived benefits. Here, we used shotgun metagenomics together with assembly and binning strategies to reconstruct metagenome-assembled genomes (MAGs) from fecal samples of a single FMT donor. We then used metagenomic mapping to track the occurrence and distribution patterns of donor MAGs in two FMT recipients. Our analyses revealed that 22% of the 92 highly complete bacterial MAGs that we identified from the donor successfully colonized and remained abundant in two recipients for at least 8 weeks. Most MAGs with a high colonization rate belonged to the order Bacteroidales. The vast majority of those that lacked evidence of colonization belonged to the order Clostridiales, and colonization success was negatively correlated with the number of genes related to sporulation. Our analysis of 151 publicly available gut metagenomes showed that the donor MAGs that colonized both recipientsmore »
- Authors:
-
- Univ. of Chicago Medicine, Chicago, IL (United States)
- Univ. of Chicago Medicine, Chicago, IL (United States); Boston Children's Hospital, Boston, MA (United States)
- Marine Biological Lab., Woods Hole, MA (United States)
- Univ. of Chicago Medicine, Chicago, IL (United States); Marine Biological Lab., Woods Hole, MA (United States)
- Publication Date:
- Research Org.:
- Argonne National Laboratory (ANL), Argonne, IL (United States)
- Sponsoring Org.:
- Gastro-Intestinal Research Foundation (GIRF); University of Chicago; USDOE
- OSTI Identifier:
- 1415981
- Grant/Contract Number:
- AC02-06CH11357
- Resource Type:
- Accepted Manuscript
- Journal Name:
- Microbiome
- Additional Journal Information:
- Journal Volume: 5; Journal Issue: 1; Journal ID: ISSN 2049-2618
- Publisher:
- BioMed Central
- Country of Publication:
- United States
- Language:
- English
- Subject:
- 60 APPLIED LIFE SCIENCES; 59 BASIC BIOLOGICAL SCIENCES; colonization; fecal microbiota transplantation; metagenome-assembled genomes; metagenomics
Citation Formats
Lee, Sonny T. M., Kahn, Stacy A., Delmont, Tom O., Shaiber, Alon, Esen, Ozcan C., Hubert, Nathaniel A., Morrison, Hilary G., Antonopoulos, Dionysios A., Rubin, David T., and Eren, A. Murat. Tracking microbial colonization in fecal microbiota transplantation experiments via genome-resolved metagenomics. United States: N. p., 2017.
Web. doi:10.1186/s40168-017-0270-x.
Lee, Sonny T. M., Kahn, Stacy A., Delmont, Tom O., Shaiber, Alon, Esen, Ozcan C., Hubert, Nathaniel A., Morrison, Hilary G., Antonopoulos, Dionysios A., Rubin, David T., & Eren, A. Murat. Tracking microbial colonization in fecal microbiota transplantation experiments via genome-resolved metagenomics. United States. https://doi.org/10.1186/s40168-017-0270-x
Lee, Sonny T. M., Kahn, Stacy A., Delmont, Tom O., Shaiber, Alon, Esen, Ozcan C., Hubert, Nathaniel A., Morrison, Hilary G., Antonopoulos, Dionysios A., Rubin, David T., and Eren, A. Murat. Thu .
"Tracking microbial colonization in fecal microbiota transplantation experiments via genome-resolved metagenomics". United States. https://doi.org/10.1186/s40168-017-0270-x. https://www.osti.gov/servlets/purl/1415981.
@article{osti_1415981,
title = {Tracking microbial colonization in fecal microbiota transplantation experiments via genome-resolved metagenomics},
author = {Lee, Sonny T. M. and Kahn, Stacy A. and Delmont, Tom O. and Shaiber, Alon and Esen, Ozcan C. and Hubert, Nathaniel A. and Morrison, Hilary G. and Antonopoulos, Dionysios A. and Rubin, David T. and Eren, A. Murat},
abstractNote = {Fecal microbiota transplantation (FMT) is an effective treatment for recurrent Clostridium difficile infection and shows promise for treating other medical conditions associated with intestinal dysbioses. However, we lack a sufficient understanding of which microbial populations successfully colonize the recipient gut, and the widely used approaches to study the microbial ecology of FMT experiments fail to provide enough resolution to identify populations that are likely responsible for FMT-derived benefits. Here, we used shotgun metagenomics together with assembly and binning strategies to reconstruct metagenome-assembled genomes (MAGs) from fecal samples of a single FMT donor. We then used metagenomic mapping to track the occurrence and distribution patterns of donor MAGs in two FMT recipients. Our analyses revealed that 22% of the 92 highly complete bacterial MAGs that we identified from the donor successfully colonized and remained abundant in two recipients for at least 8 weeks. Most MAGs with a high colonization rate belonged to the order Bacteroidales. The vast majority of those that lacked evidence of colonization belonged to the order Clostridiales, and colonization success was negatively correlated with the number of genes related to sporulation. Our analysis of 151 publicly available gut metagenomes showed that the donor MAGs that colonized both recipients were prevalent, and the ones that colonized neither were rare across the participants of the Human Microbiome Project. Although our dataset showed a link between taxonomy and the colonization ability of a given MAG, we also identified MAGs that belong to the same taxon with different colonization properties, highlighting the importance of an appropriate level of resolution to explore the functional basis of colonization and to identify targets for cultivation, hypothesis generation, and testing in model systems. Lastly, the analytical strategy adopted in our study can provide genomic insights into bacterial populations that may be critical to the efficacy of FMT due to their success in gut colonization and metabolic properties, and guide cultivation efforts to investigate mechanistic underpinnings of this procedure beyond associations.},
doi = {10.1186/s40168-017-0270-x},
journal = {Microbiome},
number = 1,
volume = 5,
place = {United States},
year = {Thu May 04 00:00:00 EDT 2017},
month = {Thu May 04 00:00:00 EDT 2017}
}
Web of Science
Figures / Tables:
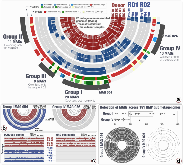 Figure 1: Distribution of MAGs across samples and HMP metagenomes. a The 92 MAGs and their level of detection in four donor samples (four inner circles) as well as two recipients (R01 and R02) before FMT (pre-FMT), 4 weeks after FMT (W4), and 8 weeks after FMT (W8). Rectangles withmore »
Figure 1: Distribution of MAGs across samples and HMP metagenomes. a The 92 MAGs and their level of detection in four donor samples (four inner circles) as well as two recipients (R01 and R02) before FMT (pre-FMT), 4 weeks after FMT (W4), and 8 weeks after FMT (W8). Rectangles withmore »
Works referenced in this record:
Anvi’o: an advanced analysis and visualization platform for ‘omics data
journal, January 2015
- Eren, A. Murat; Esen, Özcan C.; Quince, Christopher
- PeerJ, Vol. 3
Temporal and technical variability of human gut metagenomes
journal, April 2015
- Voigt, Anita Y.; Costea, Paul I.; Kultima, Jens Roat
- Genome Biology, Vol. 16, Issue 1
Long-Term Follow-Up of Colonoscopic Fecal Microbiota Transplant for Recurrent Clostridium difficile Infection
journal, January 2012
- Brandt, Lawrence J.; Aroniadis, Olga C.; Mellow, Mark
- American Journal of Gastroenterology, Vol. 107, Issue 7
Evaluation of genomic high-throughput sequencing data generated on Illumina HiSeq and Genome Analyzer systems
journal, January 2011
- Minoche, André E.; Dohm, Juliane C.; Himmelbauer, Heinz
- Genome Biology, Vol. 12, Issue 11
Systematic Review of Intestinal Microbiota Transplantation (Fecal Bacteriotherapy) for Recurrent Clostridium difficile Infection
journal, November 2011
- Gough, Ethan; Shaikh, Henna; Manges, Amee R.
- Clinical Infectious Diseases, Vol. 53, Issue 10
Fecal Transplants: What Is Being Transferred?
journal, July 2016
- Bojanova, Diana P.; Bordenstein, Seth R.
- PLOS Biology, Vol. 14, Issue 7
Review article: faecal transplantation therapy for gastrointestinal disease: Review: faecal transplantation
journal, June 2011
- Landy, J.; Al-Hassi, H. O.; McLaughlin, S. D.
- Alimentary Pharmacology & Therapeutics, Vol. 34, Issue 4
High-throughput DNA sequence analysis reveals stable engraftment of gut microbiota following transplantation of previously frozen fecal bacteria
journal, January 2013
- Hamilton, Matthew J.; Weingarden, Alexa R.; Unno, Tatsuya
- Gut Microbes, Vol. 4, Issue 2
Safety, Tolerability, and Clinical Response After Fecal Transplantation in Children and Young Adults With Ulcerative Colitis
journal, January 2013
- Kunde, Sachin; Pham, Angela; Bonczyk, Sarah
- Journal of Pediatric Gastroenterology and Nutrition, Vol. 56, Issue 6
Genome-wide analysis of the interaction between the endosymbiotic bacterium Wolbachia and its Drosophila host
journal, January 2008
- Xi, Zhiyong; Gavotte, Laurent; Xie, Yan
- BMC Genomics, Vol. 9, Issue 1
Fecal Microbiota Transplantation Induces Remission in Patients With Active Ulcerative Colitis in a Randomized Controlled Trial
journal, July 2015
- Moayyedi, Paul; Surette, Michael G.; Kim, Peter T.
- Gastroenterology, Vol. 149, Issue 1
MEGAHIT: an ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph
journal, January 2015
- Li, Dinghua; Liu, Chi-Man; Luo, Ruibang
- Bioinformatics, Vol. 31, Issue 10
Centrifuge: rapid and sensitive classification of metagenomic sequences
journal, October 2016
- Kim, Daehwan; Song, Li; Breitwieser, Florian P.
- Genome Research, Vol. 26, Issue 12
Duodenal Infusion of Donor Feces for Recurrent Clostridium difficile
journal, January 2013
- van Nood, Els; Vrieze, Anne; Nieuwdorp, Max
- New England Journal of Medicine, Vol. 368, Issue 5
Clostridium difficile infection: new developments in epidemiology and pathogenesis
journal, July 2009
- Rupnik, Maja; Wilcox, Mark H.; Gerding, Dale N.
- Nature Reviews Microbiology, Vol. 7, Issue 7
Fecal Microbiota Transplantation Inducing Remission in Crohn’s Colitis and the Associated Changes in Fecal Microbial Profile
journal, January 2014
- Kao, Dina; Hotte, Naomi; Gillevet, Patrick
- Journal of Clinical Gastroenterology, Vol. 48, Issue 7
Durable coexistence of donor and recipient strains after fecal microbiota transplantation
journal, April 2016
- Li, Simone S.; Zhu, Ana; Benes, Vladimir
- Science, Vol. 352, Issue 6285
Prodigal: prokaryotic gene recognition and translation initiation site identification
journal, March 2010
- Hyatt, Doug; Chen, Gwo-Liang; LoCascio, Philip F.
- BMC Bioinformatics, Vol. 11, Issue 1
Temporal Bacterial Community Dynamics Vary Among Ulcerative Colitis Patients After Fecal Microbiota Transplantation
journal, January 2013
- Angelberger, Sieglinde; Reinisch, Walter; Makristathis, Athanasios
- American Journal of Gastroenterology, Vol. 108, Issue 10
CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes
journal, May 2015
- Parks, Donovan H.; Imelfort, Michael; Skennerton, Connor T.
- Genome Research, Vol. 25, Issue 7
Gastrointestinal Microflora Studies in Late‐Onset Autism
journal, September 2002
- Finegold, Sydney M.; Molitoris, Denise; Song, Yuli
- Clinical Infectious Diseases, Vol. 35, Issue s1
An integrated metagenomics pipeline for strain profiling reveals novel patterns of bacterial transmission and biogeography
journal, October 2016
- Nayfach, Stephen; Rodriguez-Mueller, Beltran; Garud, Nandita
- Genome Research, Vol. 26, Issue 11
Relationship Between Thyroid Autoimmunity and Yersinia Enterocolitica Antibodies
journal, July 2002
- Çorapçioğlu, Demet; Tonyukuk, Vedia; Kiyan, Mehmet
- Thyroid, Vol. 12, Issue 7
Transfusion-transmitted infections
journal, June 2007
- Bihl, Florian; Castelli, Damiano; Marincola, Francesco
- Journal of Translational Medicine, Vol. 5, Issue 1
UGA is an additional glycine codon in uncultured SR1 bacteria from the human microbiota
journal, March 2013
- Campbell, J. H.; O'Donoghue, P.; Campbell, A. G.
- Proceedings of the National Academy of Sciences, Vol. 110, Issue 14, p. 5540-5545
A core gut microbiome in obese and lean twins
journal, November 2008
- Turnbaugh, Peter J.; Hamady, Micah; Yatsunenko, Tanya
- Nature, Vol. 457, Issue 7228
Colonization potential to reconstitute a microbe community in patients detected early after fecal microbe transplant for recurrent C. difficile
journal, January 2016
- Kumar, Ranjit; Maynard, Craig L.; Eipers, Peter
- BMC Microbiology, Vol. 16, Issue 1
Fecal Microbiota Transplantation — An Old Therapy Comes of Age
journal, January 2013
- Kelly, Ciarán P.
- New England Journal of Medicine, Vol. 368, Issue 5
Seres's pioneering microbiome drug fails mid-stage trial
journal, October 2016
- Ratner, Mark
- Nature Biotechnology, Vol. 34, Issue 10
A patient with severe Crohn's colitis responds to Faecal Microbiota Transplantation
journal, March 2014
- Gordon, Hannah; Harbord, Marcus
- Journal of Crohn's and Colitis, Vol. 8, Issue 3
Recovery of the Gut Microbiome following Fecal Microbiota Transplantation
journal, June 2014
- Seekatz, Anna M.; Aas, Johannes; Gessert, Charles E.
- mBio, Vol. 5, Issue 3
Fecal Microbiota Transplantation for Clostridium difficile Infection: Systematic Review and Meta-Analysis
journal, January 2013
- Kassam, Zain; Lee, Christine H.; Yuan, Yuhong
- American Journal of Gastroenterology, Vol. 108, Issue 4
Reset of a critically disturbed microbial ecosystem: faecal transplant in recurrent Clostridium difficile infection
journal, February 2014
- Fuentes, Susana; van Nood, Els; Tims, Sebastian
- The ISME Journal, Vol. 8, Issue 8
Systematic review: faecal microbiota transplantation in the management of inflammatory bowel disease
journal, July 2012
- Anderson, J. L.; Edney, R. J.; Whelan, K.
- Alimentary Pharmacology & Therapeutics, Vol. 36, Issue 6
Transfusion Reactions Due to Bacterial Contamination of Blood and Blood Products
journal, March 1991
- Morduchowicz, Gabriel; Pitlik, Silvio D.; Huminer, David
- Clinical Infectious Diseases, Vol. 13, Issue 2
Dynamic changes in short- and long-term bacterial composition following fecal microbiota transplantation for recurrent Clostridium difficile infection
journal, March 2015
- Weingarden, Alexa; González, Antonio; Vázquez-Baeza, Yoshiki
- Microbiome, Vol. 3, Issue 1
Systematic Review: Adverse Events of Fecal Microbiota Transplantation
journal, August 2016
- Wang, Sinan; Xu, Mengque; Wang, Weiqiang
- PLOS ONE, Vol. 11, Issue 8
Binning metagenomic contigs by coverage and composition
journal, September 2014
- Alneberg, Johannes; Bjarnason, Brynjar Smári; de Bruijn, Ino
- Nature Methods, Vol. 11, Issue 11
Environmental Genome Shotgun Sequencing of the Sargasso Sea
journal, April 2004
- Venter, J. C.
- Science, Vol. 304, Issue 5667
MetaPhlAn2 for enhanced metagenomic taxonomic profiling
journal, September 2015
- Truong, Duy Tin; Franzosa, Eric A.; Tickle, Timothy L.
- Nature Methods, Vol. 12, Issue 10
Patient-Specific Bacteroides Genome Variants in Pouchitis
journal, November 2016
- Vineis, Joseph H.; Ringus, Daina L.; Morrison, Hilary G.
- mBio, Vol. 7, Issue 6
Fast gapped-read alignment with Bowtie 2
journal, March 2012
- Langmead, Ben; Salzberg, Steven L.
- Nature Methods, Vol. 9, Issue 4
Machine Learning Meta-analysis of Large Metagenomic Datasets: Tools and Biological Insights
journal, July 2016
- Pasolli, Edoardo; Truong, Duy Tin; Malik, Faizan
- PLOS Computational Biology, Vol. 12, Issue 7
Community structure and metabolism through reconstruction of microbial genomes from the environment
journal, February 2004
- Tyson, Gene W.; Chapman, Jarrod; Hugenholtz, Philip
- Nature, Vol. 428, Issue 6978
A Novel Microbiome Therapeutic Increases Gut Microbial Diversity and Prevents Recurrent Clostridium difficile Infection
journal, February 2016
- Khanna, Sahil; Pardi, Darrell S.; Kelly, Colleen R.
- Journal of Infectious Diseases, Vol. 214, Issue 2
Should We Standardize the 1,700-Year-Old Fecal Microbiota Transplantation?
journal, January 2012
- Zhang, Faming; Luo, Wensheng; Shi, Yan
- American Journal of Gastroenterology, Vol. 107, Issue 11
PhyloSift: phylogenetic analysis of genomes and metagenomes
journal, January 2014
- Darling, Aaron E.; Jospin, Guillaume; Lowe, Eric
- PeerJ, Vol. 2
HMMER web server: interactive sequence similarity searching
journal, May 2011
- Finn, R. D.; Clements, J.; Eddy, S. R.
- Nucleic Acids Research, Vol. 39, Issue suppl
16S rRNA Gene Sequencing for Bacterial Identification in the Diagnostic Laboratory: Pluses, Perils, and Pitfalls
journal, July 2007
- Janda, J. M.; Abbott, S. L.
- Journal of Clinical Microbiology, Vol. 45, Issue 9
Insights into the phylogeny and coding potential of microbial dark matter
journal, July 2013
- Rinke, Christian; Schwientek, Patrick; Sczyrba, Alexander
- Nature, Vol. 499, Issue 7459
FastTree: Computing Large Minimum Evolution Trees with Profiles instead of a Distance Matrix
journal, April 2009
- Price, M. N.; Dehal, P. S.; Arkin, A. P.
- Molecular Biology and Evolution, Vol. 26, Issue 7
The Sequence Alignment/Map format and SAMtools
journal, June 2009
- Li, H.; Handsaker, B.; Wysoker, A.
- Bioinformatics, Vol. 25, Issue 16
A Filtering Method to Generate High Quality Short Reads Using Illumina Paired-End Technology
journal, June 2013
- Eren, A. Murat; Vineis, Joseph H.; Morrison, Hilary G.
- PLoS ONE, Vol. 8, Issue 6
Identifying biologically relevant differences between metagenomic communities
journal, February 2010
- Parks, Donovan H.; Beiko, Robert G.
- Bioinformatics, Vol. 26, Issue 6
Bowel‐flora alteration: a potential cure for inflammatory bowel disease and irritable bowel syndrome?
journal, May 1989
- Borody, T. J.; George, L.; Andrews, P.
- Medical Journal of Australia, Vol. 150, Issue 10
PhyloSift: Phylogenetic analysis of genomes and metagenomes
dataset, January 2013
- Bik, Holly; Matsen, Erick; Lowe, Eric
- figshare
PhyloSift: Phylogenetic analysis of genomes and metagenomes
dataset, January 2013
- Bik, Holly; Matsen, Erick; Lowe, Eric
- Figshare
MEGAHIT: An ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph
preprint, January 2014
- Li, Dinghua; Liu, Chi-Man; Luo, Ruibang
- arXiv
Salinimonas marina sp. nov. Isolated from Jeju Island Marine Sediment
journal, June 2021
- Kim, Minji; Lee, Ki-Eun; Cha, In-Tae
- Current Microbiology, Vol. 78, Issue 8
Fecal Microbiota Transplantation for Clostridioides difficile in High-Risk Older Adults: Treat Early, Treat Often
journal, April 2020
- Agrawal, Gaurav
- Digestive Diseases and Sciences, Vol. 65, Issue 12
A patient with severe Crohn's colitis responds to Faecal Microbiota Transplantation
journal, March 2014
- Gordon, Hannah; Harbord, Marcus
- Journal of Crohn's and Colitis, Vol. 8, Issue 3
Should We Standardize the 1,700-Year-Old Fecal Microbiota Transplantation?
journal, January 2012
- Zhang, Faming; Luo, Wensheng; Shi, Yan
- American Journal of Gastroenterology, Vol. 107, Issue 11
Long-Term Follow-Up of Colonoscopic Fecal Microbiota Transplant for Recurrent Clostridium difficile Infection
journal, January 2012
- Brandt, Lawrence J.; Aroniadis, Olga C.; Mellow, Mark
- American Journal of Gastroenterology, Vol. 107, Issue 7
Temporal Bacterial Community Dynamics Vary Among Ulcerative Colitis Patients After Fecal Microbiota Transplantation
journal, January 2013
- Angelberger, Sieglinde; Reinisch, Walter; Makristathis, Athanasios
- American Journal of Gastroenterology, Vol. 108, Issue 10
Fecal Microbiota Transplantation for Clostridium difficile Infection: Systematic Review and Meta-Analysis
journal, January 2013
- Kassam, Zain; Lee, Christine H.; Yuan, Yuhong
- American Journal of Gastroenterology, Vol. 108, Issue 4
Community structure and metabolism through reconstruction of microbial genomes from the environment
journal, February 2004
- Tyson, Gene W.; Chapman, Jarrod; Hugenholtz, Philip
- Nature, Vol. 428, Issue 6978
A core gut microbiome in obese and lean twins
journal, November 2008
- Turnbaugh, Peter J.; Hamady, Micah; Yatsunenko, Tanya
- Nature, Vol. 457, Issue 7228
Insights into the phylogeny and coding potential of microbial dark matter
journal, July 2013
- Rinke, Christian; Schwientek, Patrick; Sczyrba, Alexander
- Nature, Vol. 499, Issue 7459
Seres's pioneering microbiome drug fails mid-stage trial
journal, October 2016
- Ratner, Mark
- Nature Biotechnology, Vol. 34, Issue 10
MetaPhlAn2 for enhanced metagenomic taxonomic profiling
journal, September 2015
- Truong, Duy Tin; Franzosa, Eric A.; Tickle, Timothy L.
- Nature Methods, Vol. 12, Issue 10
Clostridium difficile infection: new developments in epidemiology and pathogenesis
journal, July 2009
- Rupnik, Maja; Wilcox, Mark H.; Gerding, Dale N.
- Nature Reviews Microbiology, Vol. 7, Issue 7
Characterizing the molecular regulation of inhibitory immune checkpoints with multimodal single-cell screens
journal, March 2021
- Papalexi, Efthymia; Mimitou, Eleni P.; Butler, Andrew W.
- Nature Genetics, Vol. 53, Issue 3
Fecal Microbiota Transplantation Induces Remission in Patients With Active Ulcerative Colitis in a Randomized Controlled Trial
journal, July 2015
- Moayyedi, Paul; Surette, Michael G.; Kim, Peter T.
- Gastroenterology, Vol. 149, Issue 1
UGA is an additional glycine codon in uncultured SR1 bacteria from the human microbiota
journal, March 2013
- Campbell, J. H.; O'Donoghue, P.; Campbell, A. G.
- Proceedings of the National Academy of Sciences, Vol. 110, Issue 14, p. 5540-5545
Gastrointestinal Microflora Studies in Late‐Onset Autism
journal, September 2002
- Finegold, Sydney M.; Molitoris, Denise; Song, Yuli
- Clinical Infectious Diseases, Vol. 35, Issue s1
MEGAHIT: an ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph
journal, January 2015
- Li, Dinghua; Liu, Chi-Man; Luo, Ruibang
- Bioinformatics, Vol. 31, Issue 10
A Novel Microbiome Therapeutic Increases Gut Microbial Diversity and Prevents Recurrent Clostridium difficile Infection
journal, February 2016
- Khanna, Sahil; Pardi, Darrell S.; Kelly, Colleen R.
- Journal of Infectious Diseases, Vol. 214, Issue 2
Is Crohn’s Disease Ready for Fecal Microbiota Transplantation?
journal, August 2014
- Borody, Thomas J.; Finlayson, Sarah; Paramsothy, Sudarshan
- Journal of Clinical Gastroenterology, Vol. 48, Issue 7
Fecal Flora Reconstitution for Recurrent Clostridium difficile Infection: Results and Methodology
journal, January 2010
- Rohlke, Faith; Surawicz, Christina M.; Stollman, Neil
- Journal of Clinical Gastroenterology, Vol. 44, Issue 8
Medical Stool
journal, June 2013
- Davidovics, Zev H.; Sylvester, Francisco A.
- Journal of Pediatric Gastroenterology & Nutrition, Vol. 56, Issue 6
An integrated metagenomics pipeline for strain profiling reveals novel patterns of bacterial transmission and biogeography
journal, October 2016
- Nayfach, Stephen; Rodriguez-Mueller, Beltran; Garud, Nandita
- Genome Research, Vol. 26, Issue 11
Systematic review: faecal microbiota transplantation in the management of inflammatory bowel disease
journal, July 2012
- Anderson, J. L.; Edney, R. J.; Whelan, K.
- Alimentary Pharmacology & Therapeutics, Vol. 36, Issue 6
Environmental Genome Shotgun Sequencing of the Sargasso Sea
journal, April 2004
- Venter, J. C.
- Science, Vol. 304, Issue 5667
Durable coexistence of donor and recipient strains after fecal microbiota transplantation
journal, April 2016
- Li, Simone S.; Zhu, Ana; Benes, Vladimir
- Science, Vol. 352, Issue 6285
Transfer of Viral Communities between Human Individuals during Fecal Microbiota Transplantation
journal, March 2016
- Chehoud, Christel; Dryga, Anatoly; Hwang, Young
- mBio, Vol. 7, Issue 2
Fecal microbiota transplantation in relapsing Clostridium difficile infection
journal, July 2012
- Rohlke, Faith; Stollman, Neil
- Therapeutic Advances in Gastroenterology, Vol. 5, Issue 6
Prodigal: prokaryotic gene recognition and translation initiation site identification
journal, March 2010
- Hyatt, Doug; Chen, Gwo-Liang; LoCascio, Philip F.
- BMC Bioinformatics, Vol. 11, Issue 1
The RAST Server: Rapid Annotations using Subsystems Technology
journal, January 2008
- Aziz, Ramy K.; Bartels, Daniela; Best, Aaron A.
- BMC Genomics, Vol. 9, Issue 1, Article No. 75
Transfusion-transmitted infections
journal, June 2007
- Bihl, Florian; Castelli, Damiano; Marincola, Francesco
- Journal of Translational Medicine, Vol. 5, Issue 1
Evaluation of genomic high-throughput sequencing data generated on Illumina HiSeq and Genome Analyzer systems
journal, January 2011
- Minoche, André E.; Dohm, Juliane C.; Himmelbauer, Heinz
- Genome Biology, Vol. 12, Issue 11
Machine Learning Meta-analysis of Large Metagenomic Datasets: Tools and Biological Insights
journal, July 2016
- Pasolli, Edoardo; Truong, Duy Tin; Malik, Faizan
- PLOS Computational Biology, Vol. 12, Issue 7
A Filtering Method to Generate High Quality Short Reads Using Illumina Paired-End Technology
journal, June 2013
- Eren, A. Murat; Vineis, Joseph H.; Morrison, Hilary G.
- PLoS ONE, Vol. 8, Issue 6
High-throughput DNA sequence analysis reveals stable engraftment of gut microbiota following transplantation of previously frozen fecal bacteria
journal, January 2013
- Hamilton, Matthew J.; Weingarden, Alexa R.; Unno, Tatsuya
- Gut Microbes, Vol. 4, Issue 2
Works referencing / citing this record:
dRep: a tool for fast and accurate genomic comparisons that enables improved genome recovery from metagenomes through de-replication
journal, July 2017
- Olm, Matthew R.; Brown, Christopher T.; Brooks, Brandon
- The ISME Journal, Vol. 11, Issue 12
Assessing the viability of transplanted gut microbiota by sequential tagging with D-amino acid-based metabolic probes
journal, March 2019
- Wang, Wei; Lin, Liyuan; Du, Yahui
- Nature Communications, Vol. 10, Issue 1
Protein-level assembly increases protein sequence recovery from metagenomic samples manyfold
journal, June 2019
- Steinegger, Martin; Mirdita, Milot; Söding, Johannes
- Nature Methods, Vol. 16, Issue 7
Elucidating Syntrophic Butyrate-Degrading Populations in Anaerobic Digesters Using Stable-Isotope-Informed Genome-Resolved Metagenomics
journal, August 2019
- Ziels, Ryan M.; Nobu, Masaru K.; Sousa, Diana Z.
- mSystems, Vol. 4, Issue 4
Long-term colonisation with donor bacteriophages following successful faecal microbial transplantation
journal, December 2018
- Draper, L. A.; Ryan, F. J.; Smith, M. K.
- Microbiome, Vol. 6, Issue 1
Sunbeam: an extensible pipeline for analyzing metagenomic sequencing experiments
journal, March 2019
- Clarke, Erik L.; Taylor, Louis J.; Zhao, Chunyu
- Microbiome, Vol. 7, Issue 1
Estimating relative biomasses of organisms in microbiota using “phylopeptidomics”
journal, March 2020
- Pible, Olivier; Allain, François; Jouffret, Virginie
- Microbiome, Vol. 8, Issue 1
Identification of the microbial diversity after fecal microbiota transplantation therapy for chronic intractable constipation using 16s rRNA amplicon sequencing
journal, March 2019
- Ohara, Tadashi
- PLOS ONE, Vol. 14, Issue 3
Sunbeam: an extensible pipeline for analyzing metagenomic sequencing experiments
posted_content, January 2019
- Clarke, Erik L.; Taylor, Louis J.; Zhao, Chunyu
Assessing the viability of transplanted gut microbiota by sequential tagging with D-amino acid-based metabolic probes
journal, March 2019
- Wang, Wei; Lin, Liyuan; Du, Yahui
- Nature Communications, Vol. 10, Issue 1
Elucidating Syntrophic Butyrate-Degrading Populations in Anaerobic Digesters Using Stable-Isotope-Informed Genome-Resolved Metagenomics
journal, August 2019
- Ziels, Ryan M.; Nobu, Masaru K.; Sousa, Diana Z.
- mSystems, Vol. 4, Issue 4
Long-term colonisation with donor bacteriophages following successful faecal microbial transplantation
journal, December 2018
- Draper, L. A.; Ryan, F. J.; Smith, M. K.
- Microbiome, Vol. 6, Issue 1
Sunbeam: an extensible pipeline for analyzing metagenomic sequencing experiments
journal, March 2019
- Clarke, Erik L.; Taylor, Louis J.; Zhao, Chunyu
- Microbiome, Vol. 7, Issue 1
Figures / Tables found in this record:

 Search WorldCat to find libraries that may hold this journal
Search WorldCat to find libraries that may hold this journal