Supersplat--spliced RNA-seq alignment
Journal Article
·
· Bioinformatics
- Oregon State Univ., Corvallis, OR (United States). Center for Genome Research and Biocomputing. Dept. of Botany and Plant Pathology; Oregon State Univ., Corvallis, OR (United States). Dept. of Electrical Engineering and Computer Science; DOE/OSTI
- Oregon State Univ., Corvallis, OR (United States). Center for Genome Research and Biocomputing. Dept. of Botany and Plant Pathology
- Oregon State Univ., Corvallis, OR (United States). Dept. of Electrical Engineering and Computer Science
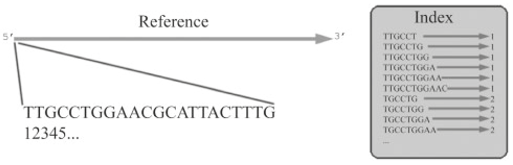
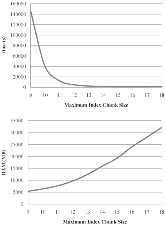
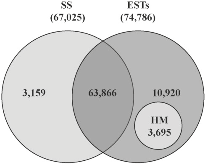
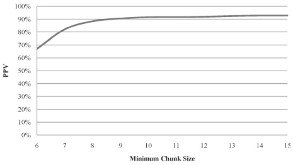
Motivation: High-throughput sequencing technologies have recently made deep interrogation of expressed transcript sequences practical, both economically and temporally. Identification of intron/exon boundaries is an essential part of genome annotation, yet remains a challenge. Here, we present supersplat, a method for unbiased splice-junction discovery through empirical RNA-seq data. Results: Using a genomic reference and RNA-seq high-throughput sequencing datasets, supersplat empirically identifies potential splice junctions at a rate of ~11.4 million reads per hour. We further benchmark the performance of the algorithm by mapping Illumina RNA-seq reads to identify introns in the genome of the reference dicot plant Arabidopsis thaliana and we demonstrate the utility of supersplat for de novo empirical annotation of splice junctions using the reference monocot plant Brachypodium distachyon. Availability: Implemented in C++, supersplat source code and binaries are freely available on the web at http://mocklerlabtools.cgrb.oregonstate.edu/
- Research Organization:
- Oregon State Univ., Corvallis, OR (United States)
- Sponsoring Organization:
- USDOE Office of Science (SC), Biological and Environmental Research (BER). Biological Systems Science Division
- Grant/Contract Number:
- FG02-08ER64630
- OSTI ID:
- 1625268
- Journal Information:
- Bioinformatics, Journal Name: Bioinformatics Journal Issue: 12 Vol. 26; ISSN 1367-4803
- Publisher:
- Oxford University PressCopyright Statement
- Country of Publication:
- United States
- Language:
- English
Similar Records
Defining the maize transcriptome de novo using deep RNA-Seq
Conference
·
Wed Jun 01 00:00:00 EDT 2011
·
OSTI ID:1050660