Consistent Metagenome-Derived Metrics Verify and Delineate Bacterial Species Boundaries
- Univ. of California, Berkeley, CA (United States)
- Univ. of California, Berkeley, CA (United States); Lawrence Berkeley National Lab. (LBNL), Berkeley, CA (United States); Chan Zuckerberg Biohub, San Francisco, CA (United States)
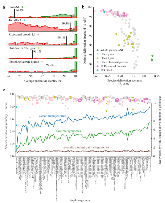
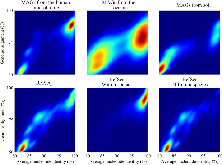
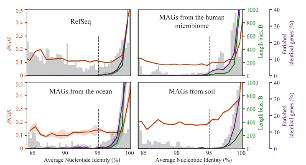
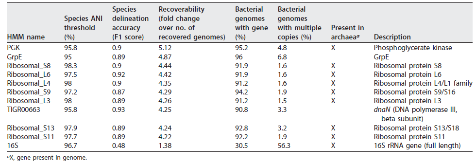
Longstanding questions relate to the existence of naturally distinct bacterial species and genetic approaches to distinguish them. Bacterial genomes in public databases form distinct groups, but these databases are subject to isolation and deposition biases. To avoid these biases, we compared 5,203 bacterial genomes from 1,457 environmental metagenomic samples to test for distinct clouds of diversity and evaluated metrics that could be used to define the species boundary. Bacterial genomes from the human gut, soil, and the ocean all exhibited gaps in whole-genome average nucleotide identities (ANI) near the previously suggested species threshold of 95% ANI. While genome-wide ratios of nonsynonymous and synonymous nucleotide differences (dN/dS) decrease until ANI values approach ~98%, two methods for estimating homologous recombination approached zero at ~95% ANI, supporting breakdown of recombination due to sequence divergence as a species-forming force. We evaluated 107 genome-based metrics for their ability to distinguish species when full genomes are not recovered. Full-length 16S rRNA genes were least useful, in part because they were underrecovered from metagenomes. However, many ribosomal proteins displayed both high metagenomic recoverability and species discrimination power. Taken together, our results verify the existence of sequence-discrete microbial species in metagenome-derived genomes and highlight the usefulness of ribosomal genes for gene-level species discrimination.
- Research Organization:
- Lawrence Berkeley National Laboratory (LBNL), Berkeley, CA (United States)
- Sponsoring Organization:
- Alfred P. Sloan Foundation; National Institutes of Health (NIH); National Science Foundation (NSF); USDOE Office of Science (SC), Biological and Environmental Research (BER)
- Grant/Contract Number:
- AC02-05CH11231
- OSTI ID:
- 1615290
- Journal Information:
- mSystems, Journal Name: mSystems Journal Issue: 1 Vol. 5; ISSN 2379-5077
- Publisher:
- American Society for MicrobiologyCopyright Statement
- Country of Publication:
- United States
- Language:
- English
Genomic diversity affects the accuracy of bacterial single-nucleotide polymorphism–calling pipelines
|
journal | February 2020 |
Similar Records
Biases in genome reconstruction from metagenomic data
Journal Article
·
Thu Oct 29 20:00:00 EDT 2020
· PeerJ
·
OSTI ID:1693794