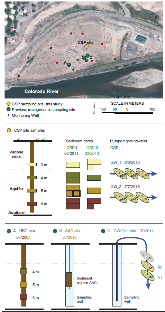
Aquifer environment selects for microbial species cohorts in sediment and groundwater
- Univ. of California, Berkeley, CA (United States). Dept. of Earth and Planetary Science
- Univ. of California, Berkeley, CA (United States). Dept. of Plant and Microbial Biology
- Columbia Univ., New York, NY (United States). Dept. of Earth and Environmental Science
- Lawrence Berkeley National Lab. (LBNL), Berkeley, CA (United States). Geophysics Dept. Earth Sciences Division
- USDOE Joint Genome Institute (JGI), Walnut Creek, CA (United States). Metagenome Program
- Univ. of California, Berkeley, CA (United States). Dept. of Earth and Planetary Science. Dept. of Environmental Science, Policy, and Management
Little is known about the biogeography or stability of sediment-associated microbial community membership because these environments are biologically complex and generally difficult to sample. High-throughput-sequencing methods provide new opportunities to simultaneously genomically sample and track microbial community members across a large number of sampling sites or times, with higher taxonomic resolution than is associated with 16 S ribosomal RNA gene surveys, and without the disadvantages of primer bias and gene copy number uncertainty. We characterized here a sediment community at 5 m depth in an aquifer adjacent to the Colorado River and tracked its most abundant 133 organisms across 36 different sediment and groundwater samples. We sampled sites separated by centimeters, meters and tens of meters, collected on seven occasions over 6 years. Analysis of 1.4 terabase pairs of DNA sequence showed that these 133 organisms were more consistently detected in saturated sediments than in samples from the vadose zone, from distant locations or from groundwater filtrates. Abundance profiles across aquifer locations and from different sampling times identified organism cohorts that comprised subsets of the 133 organisms that were consistently associated. The data suggest that cohorts are partly selected for by shared environmental adaptation.
- Research Organization:
- Lawrence Berkeley National Laboratory (LBNL), Berkeley, CA (United States)
- Sponsoring Organization:
- USDOE Office of Science (SC), Biological and Environmental Research (BER) (SC-23)
- Grant/Contract Number:
- AC02-05CH11231; SC0004918
- OSTI ID:
- 1512234
- Journal Information:
- The ISME Journal, Journal Name: The ISME Journal Journal Issue: 8 Vol. 9; ISSN 1751-7362
- Publisher:
- Nature Publishing GroupCopyright Statement
- Country of Publication:
- United States
- Language:
- English
The selective pressures on the microbial community in a metal-contaminated aquifer
|
journal | December 2018 |
Critical biogeochemical functions in the subsurface are associated with bacteria from new phyla and little studied lineages: N- and C-cycling organisms in the subsurface
|
journal | July 2015 |
Evidence for Microbial Mediated NO3− Cycling Within Floodplain Sediments During Groundwater Fluctuations
|
journal | July 2019 |
Similar Records
Snowmelt Induced Hydrologic Perturbations Drive Dynamic Microbiological and Geochemical Behaviors across a Shallow Riparian Aquifer
Snowmelt Induced Hydrologic Perturbations Drive Dynamic Microbiological and Geochemical Behaviors across a Shallow Riparian Aquifer
Journal Article
·
Tue May 10 20:00:00 EDT 2016
· Frontiers in Earth Science
·
OSTI ID:1425411
Snowmelt Induced Hydrologic Perturbations Drive Dynamic Microbiological and Geochemical Behaviors across a Shallow Riparian Aquifer
Journal Article
·
Wed May 11 00:00:00 EDT 2016
· Frontiers in Earth Science
·
OSTI ID:1342300