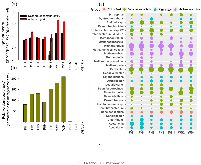
Unraveling Microbial Communities Associated with Methylmercury Production in Paddy Soils
Journal Article
·
· Environmental Science and Technology
- Huazhong Agricultural University, Wuhan (China); Chinese Academy of Sciences (CAS), Beijing (China)
- Oak Ridge National Lab. (ORNL), Oak Ridge, TN (United States)
- Chinese Academy of Sciences (CAS), Beijing (China)
- The University of Melbourne, Victoria (Australia)
- Zhejiang Univ. (China)
- Chinese Academy of Sciences (CAS), Beijing (China); The University of Melbourne, Victoria (Australia)
- Oak Ridge National Lab. (ORNL), Oak Ridge, TN (United States); Univ. of Tennessee, Knoxville, TN (United States)
Here, we report rice consumption is now recognized as an important pathway of human exposure to the neurotoxin methylmercury (MeHg), particularly in countries where rice is a staple food. Although the discovery of a two-gene cluster hgcAB has linked Hg methylation to several phylogenetically diverse groups of anaerobic microorganisms converting inorganic mercury (Hg) to MeHg, the prevalence and diversity of Hg methylators in microbial communities of rice paddy soils remain unclear. We characterized the abundance and distribution of hgcAB genes using third-generation PacBio long-read sequencing and Illumina short-read metagenomic sequencing, in combination with quantitative PCR analyses in several mine-impacted paddy soils from southwest China. Both Illumina and PacBio sequencing analyses revealed that Hg methylating communities were dominated by iron-reducing bacteria (i.e., Geobacter) and methanogens, with a relatively low abundance of hgcA+ sulfate-reducing bacteria in the soil. A positive correlation was observed between the MeHg content in soil and the relative abundance of Geobacter carrying the hgcA gene. Phylogenetic analysis also uncovered some hgcAB sequences closely related to three novel Hg methylators, Geobacter anodireducens, Desulfuromonas sp. DDH964, and Desulfovibrio sp. J2, among which G. anodireducens was validated for its ability to methylate Hg. Lastly, these findings shed new light on microbial community composition and major clades likely driving Hg methylation in rice paddy soils.
- Research Organization:
- Oak Ridge National Laboratory (ORNL), Oak Ridge, TN (United States)
- Sponsoring Organization:
- USDOE Office of Science (SC), Biological and Environmental Research (BER) (SC-23)
- Grant/Contract Number:
- AC05-00OR22725
- OSTI ID:
- 1490597
- Journal Information:
- Environmental Science and Technology, Journal Name: Environmental Science and Technology Journal Issue: 22 Vol. 52; ISSN 0013-936X
- Publisher:
- American Chemical Society (ACS)Copyright Statement
- Country of Publication:
- United States
- Language:
- English
An Improved hgcAB Primer Set and Direct High-Throughput Sequencing Expand Hg-Methylator Diversity in Nature
|
journal | October 2020 |
Biotic formation of methylmercury: A bio–physico–chemical conundrum
|
journal | November 2019 |
Intestinal Methylation and Demethylation of Mercury
|
journal | December 2018 |
Similar Records
Determining the Reliability of Measuring Mercury Cycling Gene Abundance with Correlations with Mercury and Methylmercury Concentrations
Journal Article
·
Sun Jun 30 20:00:00 EDT 2019
· Environmental Science and Technology
·
OSTI ID:1557487